Vision-based Deep Learning Analysis of Unordered Biomedical Tabular Datasets via Optimal Spatial Cartography
arXiv cs.LG / 3/25/2026
💬 OpinionIdeas & Deep AnalysisModels & Research
Key Points
- The paper addresses a key limitation in biomedical tabular modeling: features are inherently unordered, so vision-style architectures can’t directly exploit local structure and higher-order interactions.
- It introduces Dynamic Feature Mapping (Dynomap), an end-to-end differentiable framework that learns an optimal, task-driven spatial topology of tabular features via fully differentiable rendering.
- Dynomap converts high-dimensional tabular vectors into learned feature maps, enabling vision-based deep learning to work effectively on unordered biomedical inputs without heuristics or external priors.
- Experiments on multiple clinical and biological datasets show consistent improvements over classical ML, modern deep tabular models, and existing vector-to-image approaches.
- In liquid biopsy data, Dynomap improved multiclass cancer subtype prediction accuracy by up to 18% and produced coherent spatial organization of clinically relevant gene signatures, with up to 8% gains on a Parkinson disease voice dataset.
Related Articles
CRM Development That Drives Growth
Dev.to
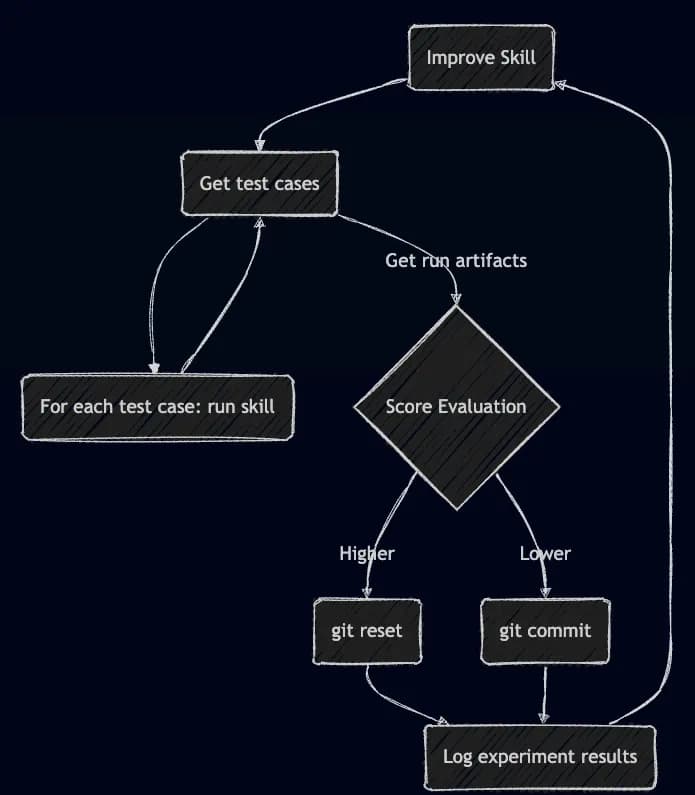
Karpathy's Autoresearch: Improving Agentic Coding Skills
Dev.to

How to Write AI Prompts That Actually Work
Dev.to
[D] Any other PhD students feel underprepared and that the bar is too low?
Reddit r/MachineLearning
Automating the Perfect Pitch: An AI Framework for Boutique PR
Dev.to