Dino-NestedUNet: Unlocking Foundation Vision Encoders for Pathology Tumor Bulk Segmentation via Dense Decoding
arXiv cs.CV / 5/5/2026
📰 NewsModels & Research
Key Points
- The paper introduces Dino-NestedUNet, which pairs a pre-trained DINOv3 vision foundation encoder with a “Nested Dense Decoder” designed for more accurate boundary reconstruction in pathology tumor bulk segmentation.
- It argues that prior approaches that freeze VFMs and attach lightweight decoders suffer from capacity mismatch, leading to poorer boundary fidelity for infiltrative tumors.
- Dino-NestedUNet replaces sparse skip connections and simple upsampling with a dense grid of intermediate pathways to support continuous feature reuse and multi-scale recalibration during decoding.
- Experiments on three histopathology cohorts (CHTN, OSU, CAMELYON16) show consistent gains over UNet++ and standard Dino-UNet variants, with especially strong benefits under cross-domain shift.
- The model also demonstrates promising external generalization via zero-shot testing (train on CHTN, test on TIGER WSIBULK and OSU CRC) without fine-tuning, highlighting the value of dense decoding for foundation-encoder segmentation tasks.
Related Articles

Singapore's Fraud Frontier: Why AI Scam Detection Demands Regulatory Precision
Dev.to

From OOM to 262K Context: Running Qwen3-Coder 30B Locally on 8GB VRAM
Dev.to

Nano Banana Pro vs DALL-E 3 vs Midjourney: A Practical Comparison From Someone Who Actually Uses All Three
Dev.to
LLMs edited 86 human essays toward a semantic cluster not occupied by any human writer [D]
Reddit r/MachineLearning
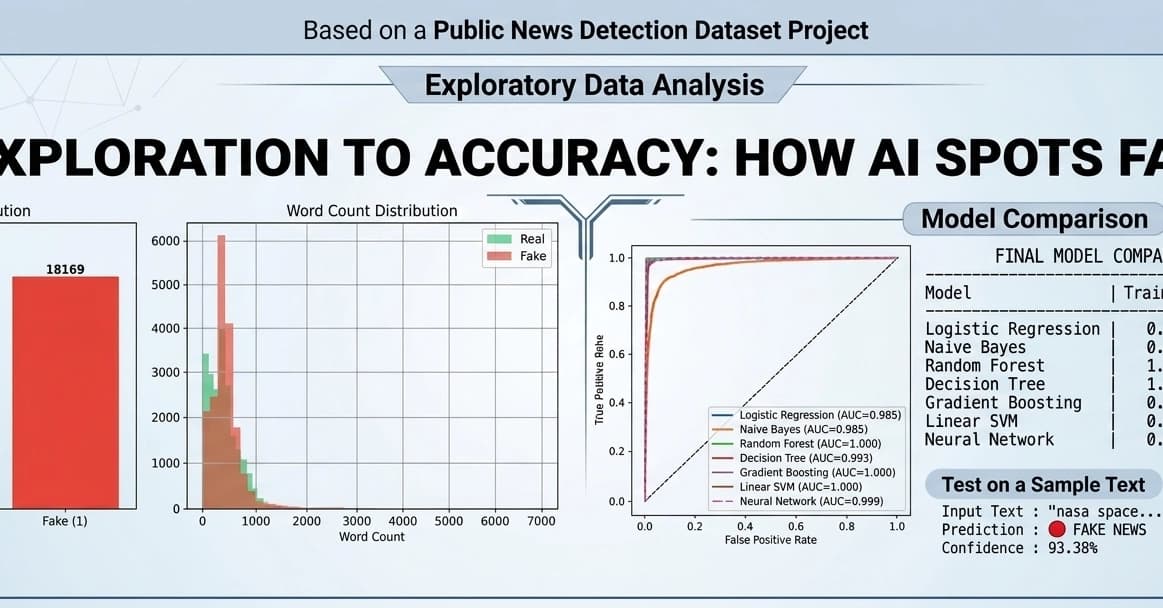
Fake News Detection using Machine Learning & NLP!
Dev.to