A Comparative Study of QSPR Methods on a Unique Multitask PAMPA dataset
arXiv cs.LG / 5/4/2026
📰 NewsIdeas & Deep AnalysisModels & Research
Key Points
- The paper introduces a new multitask PAMPA dataset covering 143 molecules tested in vitro for passive membrane permeability across six different model membranes.
- It compares multiple molecular descriptor sets and regression approaches, from simple linear regression to a pre-trained transformer model.
- The study focuses on how predictive performance must be balanced against model interpretability, especially when using machine learning methods.
- Results suggest that expert-designed physico-chemical descriptors can be more effective than deep learning representations for studies with limited sample sizes.
- The authors position the work as the most comprehensive attempt to simultaneously model multiple organ-specific PAMPA membranes so far, providing new membrane-specific permeability insights.
Related Articles
A very basic litmus test for LLMs "ok give me a python program that reads my c: and put names and folders in a sorted list from biggest to small"
Reddit r/LocalLLaMA
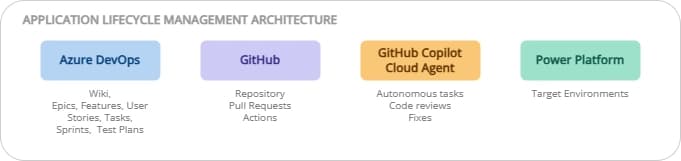
ALM on Power Platform: ADO + GitHub, the best of both worlds
Dev.to

Experiment: Does repeated usage influence ChatGPT 5.4 outputs in a RAG-like setup?
Dev.to

Find 12 high-volume, low-competition GEO content topics Topify.ai should rank on
Dev.to

When a memorized rule fits your bug too well: a meta-trap of agent workflows
Dev.to