Align Your Structures: Generating Trajectories with Structure Pretraining for Molecular Dynamics
arXiv cs.LG / 4/7/2026
📰 NewsIdeas & Deep AnalysisModels & Research
Key Points
- The paper proposes a framework for generating molecular dynamics (MD) trajectories with deep generative models by using structure pretraining to address limited MD trajectory data and the complexity of high-dimensional MD distributions.
- It trains a diffusion-based structure generation model on large-scale conformer datasets and adds an interpolator module trained on MD trajectory data to enforce temporal consistency across generated structures.
- The method decomposes MD trajectory generation into two more manageable subproblems—structural generation and temporal alignment—by leveraging abundant structural information while using MD data specifically for temporal constraints.
- Experiments on QM9 and DRUGS evaluate unconditional generation, forward simulation, and interpolation, showing improvements across geometric, dynamical, and energetic accuracy metrics.
- The framework is further extended to tetrapeptide and protein monomer systems, indicating broader applicability beyond small molecules.
Related Articles

Big Tech firms are accelerating AI investments and integration, while regulators and companies focus on safety and responsible adoption.
Dev.to

Could it be that this take is not too far fetched?
Reddit r/LocalLLaMA

npm audit Is Broken — Here's the Claude Code Skill I Built to Fix It
Dev.to

Meta Launches Muse Spark: A New AI Model for Everyday Use
Dev.to
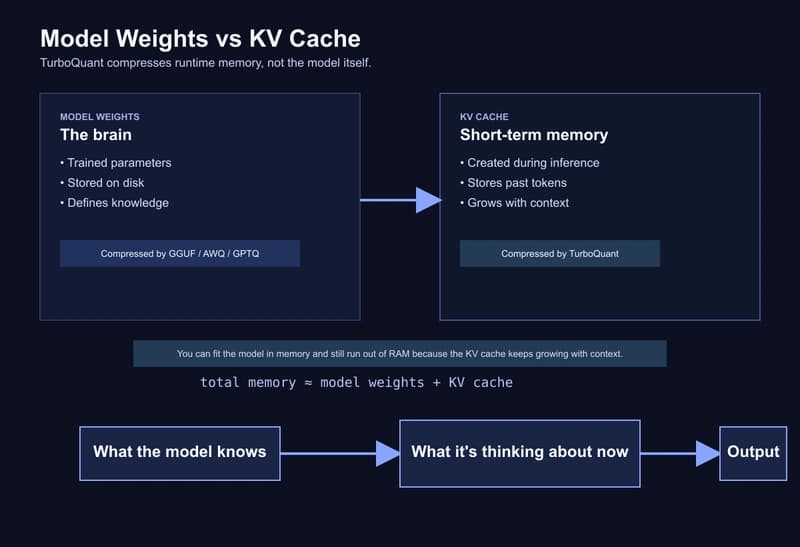
TurboQuant on a MacBook: building a one-command local stack with Ollama, MLX, and an automatic routing proxy
Dev.to