A Utility-preserving De-identification Pipeline for Cross-hospital Radiology Data Sharing
arXiv cs.CV / 4/9/2026
💬 OpinionIdeas & Deep AnalysisModels & Research
Key Points
- The paper presents a utility-preserving de-identification pipeline (UPDP) aimed at enabling cross-hospital radiology data sharing without losing clinically important signal for training medical AI models.
- UPDP uses a blacklist of privacy-sensitive terms plus a whitelist of pathology-related terms, and generates privacy-filtered but pathology-reserved synthetic radiology image counterparts.
- The approach also involves ID-filtered reports, allowing the resulting de-identified images and text to be securely shared across hospitals for downstream model development and evaluation.
- Experiments on public chest X-ray benchmarks show strong privacy removal of identity-related information while maintaining competitive diagnostic accuracy, though identity-related accuracy declines.
- In cross-hospital experiments, combining de-identified shared data with local hospital data improves performance relative to using local data alone.
Related Articles

Big Tech firms are accelerating AI investments and integration, while regulators and companies focus on safety and responsible adoption.
Dev.to

Could it be that this take is not too far fetched?
Reddit r/LocalLLaMA

npm audit Is Broken — Here's the Claude Code Skill I Built to Fix It
Dev.to

Meta Launches Muse Spark: A New AI Model for Everyday Use
Dev.to
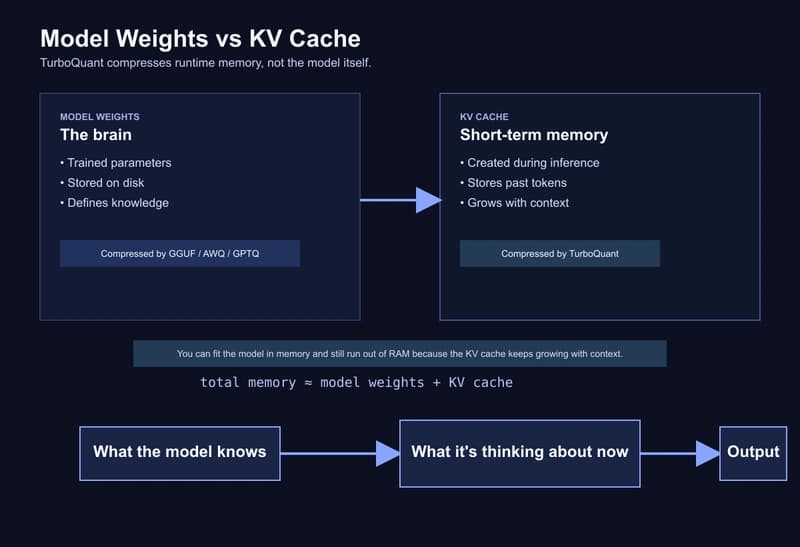
TurboQuant on a MacBook: building a one-command local stack with Ollama, MLX, and an automatic routing proxy
Dev.to