Interpretable DNA Sequence Classification via Dynamic Feature Generation in Decision Trees
arXiv cs.AI / 4/15/2026
💬 OpinionIdeas & Deep AnalysisModels & Research
Key Points
- The paper argues that while deep neural networks excel at DNA sequence prediction, they are often black boxes, motivating more interpretable approaches such as axis-aligned decision trees.
- It identifies a key limitation of standard decision-tree splitting—treating raw features independently at each node—leading to overly deep trees that hurt interpretability and generalization.
- The proposed DEFT framework adaptively generates higher-level, biologically informed sequence features during decision-tree construction rather than relying only on raw inputs.
- DEFT uses large language models to propose candidate features based on the local sequence distribution at each node and then iteratively refines them using a reflection mechanism.
- Experiments across multiple genomic tasks show that DEFT can produce human-interpretable features while maintaining strong predictive performance.
Related Articles
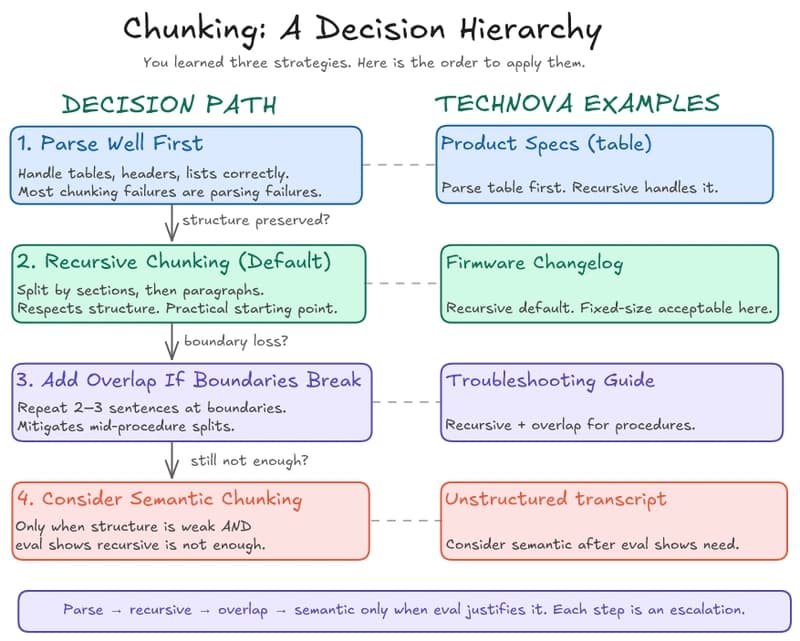
RAG in Practice — Part 4: Chunking, Retrieval, and the Decisions That Break RAG
Dev.to
Why dynamically routing multi-timescale advantages in PPO causes policy collapse (and a simple decoupled fix) [R]
Reddit r/MachineLearning

How AI Interview Assistants Are Changing Job Preparation in 2026
Dev.to

Consciousness in Artificial Intelligence: Insights from the Science ofConsciousness
Dev.to

NEW PROMPT INJECTION
Dev.to