Deep neural networks with Fisher vector encoding for medical image classification
arXiv cs.CV / 5/5/2026
📰 NewsModels & Research
Key Points
- The paper proposes using Fisher Vector (an orderless encoding method) to enhance hybrid CNN + Vision Transformer (ViT) architectures for medical image classification, especially when data availability varies.
- The approach computes Fisher Vectors by estimating a Gaussian Mixture Model (GMM) over image feature representations, aiming to improve feature representation beyond existing CNN+ViT hybrids.
- To address scalability limits on large datasets, the authors introduce a strategy that limits the growth in computational cost of GMM estimation as dataset size increases.
- Experiments on multiple medical imaging benchmarks (MedMNIST v2, Clean-CC-CCII, and ISIC2018) show improved results, outperforming benchmarks across all MedMNIST v2 datasets and achieving competitive performance on Clean-CC-CCII and ISIC2018.
- Overall, the work positions Fisher Vector-enhanced CNN+ViT models as a practical option for both small and large medical imaging datasets while managing computational trade-offs.
Related Articles

Singapore's Fraud Frontier: Why AI Scam Detection Demands Regulatory Precision
Dev.to

From OOM to 262K Context: Running Qwen3-Coder 30B Locally on 8GB VRAM
Dev.to

Nano Banana Pro vs DALL-E 3 vs Midjourney: A Practical Comparison From Someone Who Actually Uses All Three
Dev.to
LLMs edited 86 human essays toward a semantic cluster not occupied by any human writer [D]
Reddit r/MachineLearning
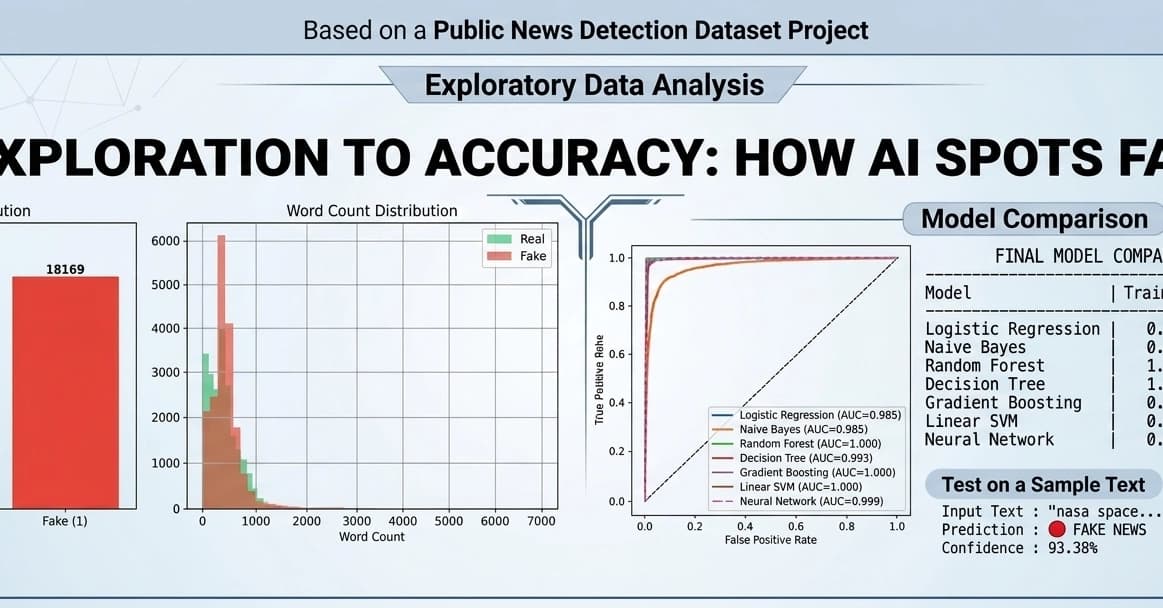
Fake News Detection using Machine Learning & NLP!
Dev.to