TriFit: Trimodal Fusion with Protein Dynamics for Mutation Fitness Prediction
arXiv cs.LG / 4/15/2026
📰 NewsSignals & Early TrendsIdeas & Deep AnalysisModels & Research
Key Points
- TriFit is presented as a multimodal supervised framework for single amino-acid variant (SAV) mutation fitness prediction that explicitly incorporates protein dynamics alongside sequence and structure.
- The model combines three embedding sources—ESM-2-based sequence embeddings, AlphaFold2-derived structural geometry embeddings, and Gaussian Network Model (GNM) dynamics features—fused via a four-expert Mixture-of-Experts (MoE) with trimodal cross-modal contrastive learning.
- TriFit adaptively learns how to weight different modality combinations per protein using an MoE router, avoiding fixed assumptions about which modality matters most.
- On the ProteinGym substitution benchmark (217 DMS assays, 696k SAVs), TriFit reports AUROC of 0.897 ± 0.0002, surpassing prior supervised baselines and improving over the best listed zero-shot model.
- Ablations indicate dynamics contributes the most additional gain beyond pairwise fusion, and the method produces well-calibrated probabilistic outputs without post-hoc calibration.
Related Articles

Black Hat Asia
AI Business

The Complete Guide to Better Meeting Productivity with AI Note-Taking
Dev.to

5 Ways Real-Time AI Can Boost Your Sales Call Performance
Dev.to
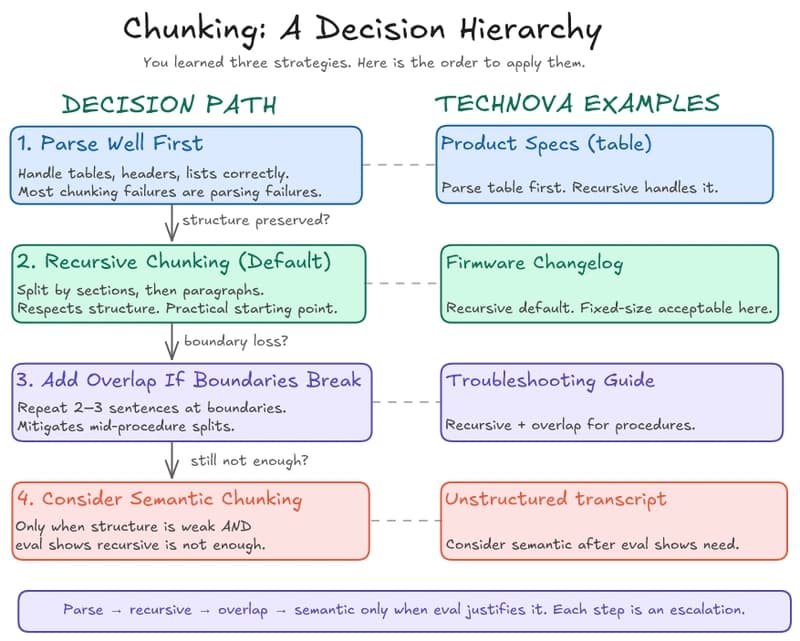
RAG in Practice — Part 4: Chunking, Retrieval, and the Decisions That Break RAG
Dev.to
Why dynamically routing multi-timescale advantages in PPO causes policy collapse (and a simple decoupled fix) [R]
Reddit r/MachineLearning