Polyformer: a generative framework for thermodynamic modeling of polymeric molecules
arXiv cs.LG / 4/17/2026
💬 OpinionModels & Research
Key Points
- Polyformer is introduced as a generative framework that performs thermodynamic modeling for polymeric molecules rather than predicting a single static structure.
- By taking a molecule’s sequence and temperature (or another thermodynamic variable) as inputs, Polyformer generates conformations that match the molecule’s thermodynamic conformational ensemble.
- The approach is positioned as the first generative model addressing folding, ensemble formation, and ensemble changes with temperature in a unified way.
- In tests on protein domains of 50–111 residues, Polyformer’s predictions show good agreement with Molecular Dynamics (MD) trajectories.
- The work extends the structural-biology paradigm from “sequence → best conformation” to “sequence + thermodynamics → ensemble behavior,” supporting more realistic modeling of biomolecular function.
Related Articles
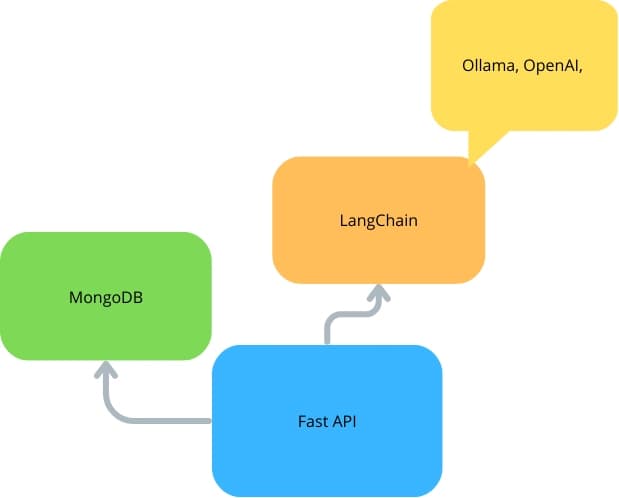
FastAPI With LangChain and MongoDB
Dev.to
![[2026] OpenTelemetry for LLM Observability — Self-Hosted Setup](/_next/image?url=https%3A%2F%2Fmedia2.dev.to%2Fdynamic%2Fimage%2Fwidth%3D1200%2Cheight%3D627%2Cfit%3Dcover%2Cgravity%3Dauto%2Cformat%3Dauto%2Fhttps%253A%252F%252Fdev-to-uploads.s3.amazonaws.com%252Fuploads%252Farticles%252Flu4b6ttuhur71z5gemm0.png&w=3840&q=75)
[2026] OpenTelemetry for LLM Observability — Self-Hosted Setup
Dev.to

The AI Education Product on Product Hunt Worth Watching
Dev.to

The joy and pain of training an LLM from scratch
Reddit r/LocalLLaMA

Did you know that you can use Qwen3.5-35B-A3B-Base as an instruction/reasoning Model?
Reddit r/LocalLLaMA