CoDaS: AI Co-Data-Scientist for Biomarker Discovery via Wearable Sensors
arXiv cs.AI / 4/17/2026
📰 NewsDeveloper Stack & InfrastructureSignals & Early TrendsModels & Research
Key Points
- The paper introduces CoDaS, an AI multi-agent system that turns continuous wearable sensor signals into clinically actionable biomarkers through iterative hypothesis generation and statistical/literature-grounded reasoning with human oversight.
- Using 9,279 participant-observations across three cohorts, CoDaS produced 41 mental-health and 25 metabolic candidate digital biomarkers, validated via replication, stability, robustness, and discriminative power checks.
- In two independent depression cohorts, CoDaS consistently identified features related to circadian instability, including sleep-duration variability and sleep-onset variability, both showing statistically significant correlations.
- For metabolic outcomes, CoDaS derived a cardiovascular fitness index and recovered established clinical associations such as AST/ALT with insulin resistance.
- Adding CoDaS-derived features to demographic variables yielded modest but reliable predictive gains (cross-validated ΔR² increases of 0.040 for depression and 0.021 for insulin resistance), supporting traceable, systematic biomarker discovery from wearable data.
Related Articles

Reported ban on ‘sex robots’ by online platform fuels debate on AI boundaries and content moderation
Reddit r/artificial
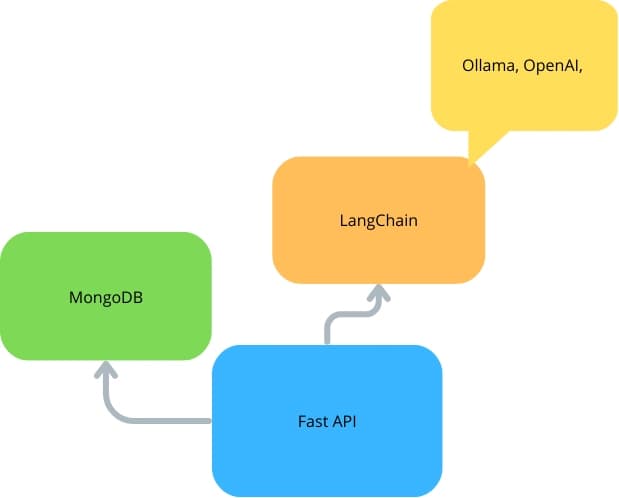
FastAPI With LangChain and MongoDB
Dev.to
Best AI Game Creator in 2026
Dev.to
![[Patterns] AI Agent Error Handling That Actually Works](/_next/image?url=https%3A%2F%2Fmedia2.dev.to%2Fdynamic%2Fimage%2Fwidth%3D1200%2Cheight%3D627%2Cfit%3Dcover%2Cgravity%3Dauto%2Cformat%3Dauto%2Fhttps%253A%252F%252Fdev-to-uploads.s3.amazonaws.com%252Fuploads%252Farticles%252Frn5czaopq2vzo7cglady.png&w=3840&q=75)
[Patterns] AI Agent Error Handling That Actually Works
Dev.to
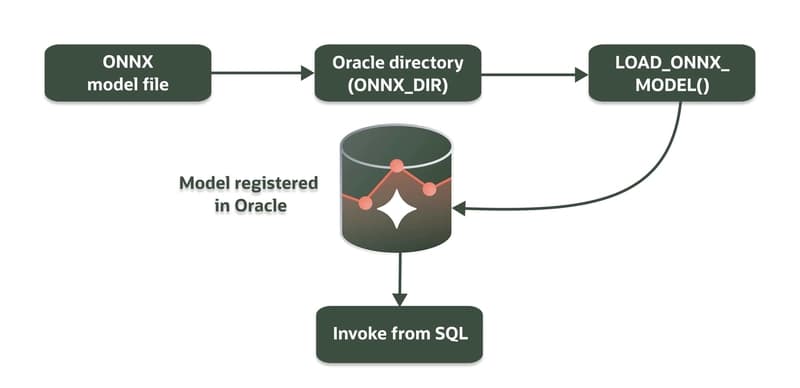
Building ONNX Embedding Workflows in Oracle AI Database with Python
Dev.to