EvoFlows: Evolutionary Edit-Based Flow-Matching for Protein Engineering
arXiv cs.LG / 3/13/2026
📰 NewsIdeas & Deep AnalysisModels & Research
Key Points
- EvoFlows is a variable-length sequence-to-sequence protein modeling approach designed for protein engineering, enabling a limited, controllable number of insertions, deletions, and substitutions on a template protein sequence.
- Unlike autoregressive and masked language models, EvoFlows predict both which mutation to perform and where it should occur by learning mutational trajectories between evolutionarily related sequences using edit flows.
- In silico evaluations on diverse protein communities from UNIREF and OAS show EvoFlows capture protein sequence distributions with quality comparable to leading masked language models while better generating non-trivial yet natural-like mutants from a given template.
- The work points to a more controllable, trajectory-aware approach to protein design that could influence future workflows and tooling in protein engineering.
Related Articles
Day 10: 230 Sessions of Hustle and It Comes Down to One Person Reading a Document
Dev.to
5 Dangerous Lies Behind Viral AI Coding Demos That Break in Production
Dev.to
Two bots, one confused server: what Nimbus revealed about AI agent identity
Dev.to
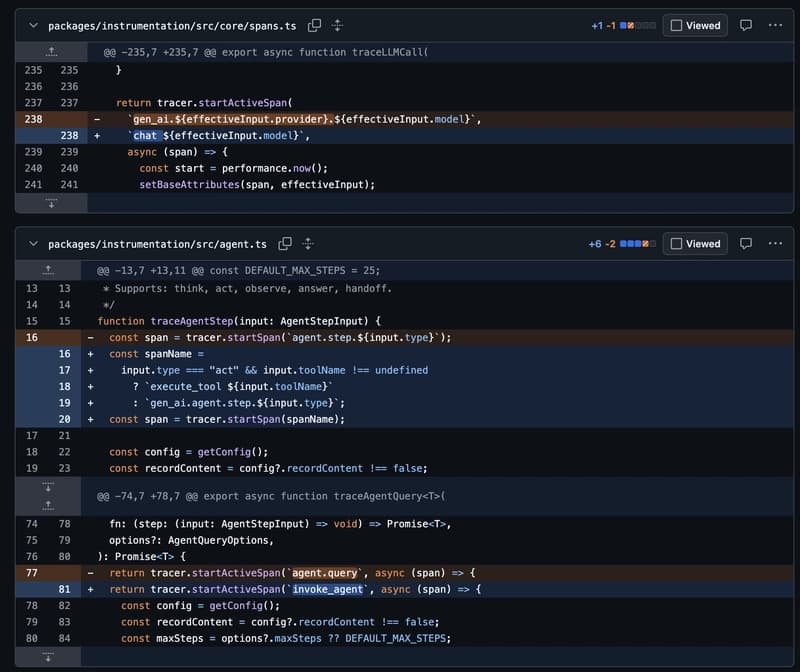
OpenTelemetry just standardized LLM tracing. Here's what it actually looks like in code.
Dev.to
PIXIU: A Large Language Model, Instruction Data and Evaluation Benchmark forFinance
Dev.to