Multi-RF Fusion with Multi-GNN Blending for Molecular Property Prediction
arXiv cs.AI / 3/24/2026
📰 NewsSignals & Early TrendsIdeas & Deep AnalysisModels & Research
Key Points
- The arXiv paper proposes Multi-RF Fusion with Multi-GNN Blending for molecular property prediction, reporting a test ROC-AUC of 0.8476 ± 0.0002 on the ogbg-molhiv benchmark (10 seeds) and claiming #1 on the OGB leaderboard over HyperFusion.
- The approach combines a rank-averaged ensemble of 12 Random Forests trained on a large concatenated fingerprint vector (FCFP, ECFP, MACCS, and atom pairs; 4,263 dimensions) with blended deep-ensembled GNN predictions at 12% weight.
- Two key improvements are highlighted: using max_features = 0.20 for Random Forests (instead of the default sqrt(d)) boosts AUC by about +0.008 on a scaffold split.
- The method reduces GNN-related randomness by averaging GNN outputs across 10 seeds before blending, effectively eliminating GNN seed variance and reducing final performance standard deviation from 0.0008 to 0.0002.
- The results are achieved without external data or any pre-training, emphasizing the effectiveness of the ensemble/blending and tuning strategy.
Related Articles
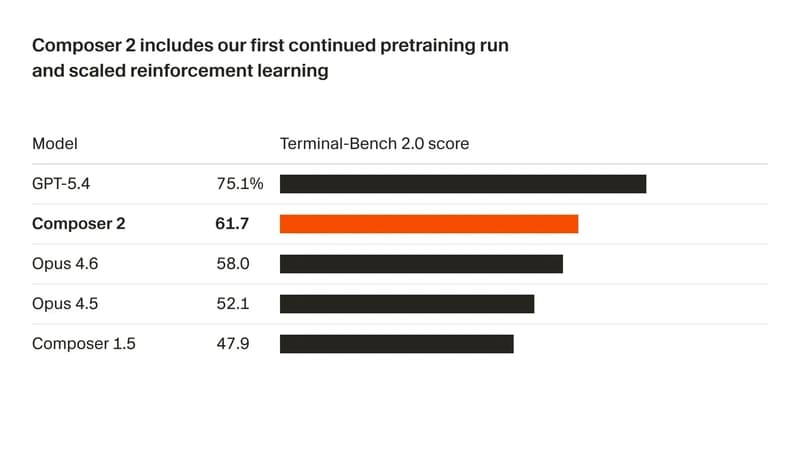
Composer 2: What is new and Compares with Claude Opus 4.6 & GPT-5.4
Dev.to
How UCP Breaks Your E-Commerce Tracking Stack: A Platform-by-Platform Analysis
Dev.to
AI Text Analyzer vs Asking Friends: Which Gives Better Perspective?
Dev.to
[D] Cathie wood claims ai productivity wave is starting, data shows 43% of ceos save 8+ hours weekly
Reddit r/MachineLearning

Microsoft hires top AI researchers from Allen Institute for AI for Suleyman's Superintelligence team
THE DECODER