Mining Negative Sequential Patterns to Improve Viral Genomic Feature Representation and Classification
arXiv cs.LG / 4/30/2026
💬 OpinionIdeas & Deep AnalysisModels & Research
Key Points
- The paper addresses the limits of existing viral genome classifiers that rely mainly on composition or frequency features, which can reduce interpretability and accuracy on complex or imbalanced datasets.
- It introduces GeneNSPCla, a viral classification framework that uses Negative Sequential Patterns (NSPs) to extract discriminative absence-based signals from RNA viral genomic sequences and converts them into feature vectors for multiple supervised classifiers.
- The authors propose GONPM+, a genomic-adapted negative pattern mining algorithm designed to find longer, more biologically meaningful negative sequential patterns.
- Experiments across 8 classifiers show that GONPM+ improves average accuracy by 10.03% over the original negative pattern mining method and by 24.75% over positive pattern mining.
- Overall, the results suggest that incorporating absence-based sequential information provides a complementary and effective perspective for viral genome representation and classification.
- .
Related Articles
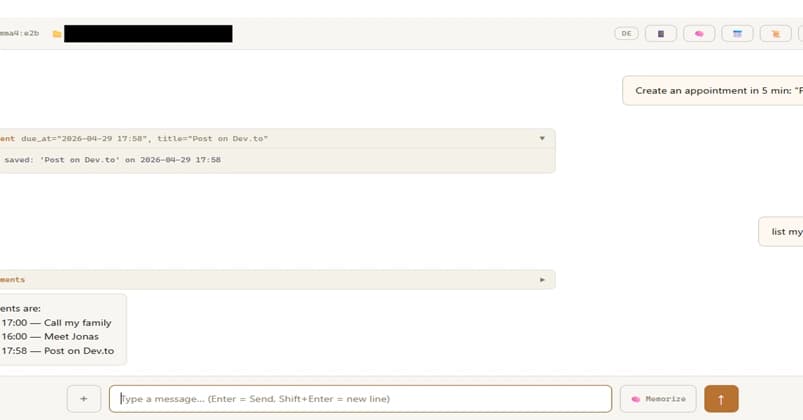
Building a Local AI Agent (Part 2): Six UX and UI Design Challenges
Dev.to

We Built a DNS-Based Discovery Protocol for AI Agents — Here's How It Works
Dev.to

Your first business opportunity in 3 commands: /register_directory in @biznode_bot, wait for matches, then /my_pulse to view...
Dev.to

Building AI Evaluation Pipelines: Automating LLM Testing from Dataset to CI/CD
Dev.to
Function Calling Harness 2: CoT Compliance from 9.91% to 100%
Dev.to