Beyond Expression Similarity: Contrastive Learning Recovers Functional Gene Associations from Protein Interaction Structure
arXiv cs.LG / 3/24/2026
💬 OpinionSignals & Early TrendsIdeas & Deep AnalysisModels & Research
Key Points
- The paper introduces a contrastive learning approach (Contrastive Association Learning, CAL) under the PAM framework, arguing that useful links arise from shared co-occurrence contexts rather than embedding similarity.
- Experiments in molecular biology show that training CAL on protein–protein interaction data recovers gene functional associations far better than gene-expression similarity, achieving cross-boundary AUCs of 0.908 (CRISPRi/K562) and 0.947 (DepMap).
- Cross-domain testing indicates inductive transfer works better in biology than in text, with node-disjoint splits yielding AUC 0.826 (+0.127 vs baselines), suggesting physically grounded interaction signals generalize.
- The authors find CAL scores anti-correlate with protein interaction degree (Spearman r = -0.590) and that improvements concentrate on understudied genes with focused interaction profiles.
- They observe that higher-quality association data can outperform larger but noisier training sets, with results stable across random seeds and threshold choices.
Related Articles
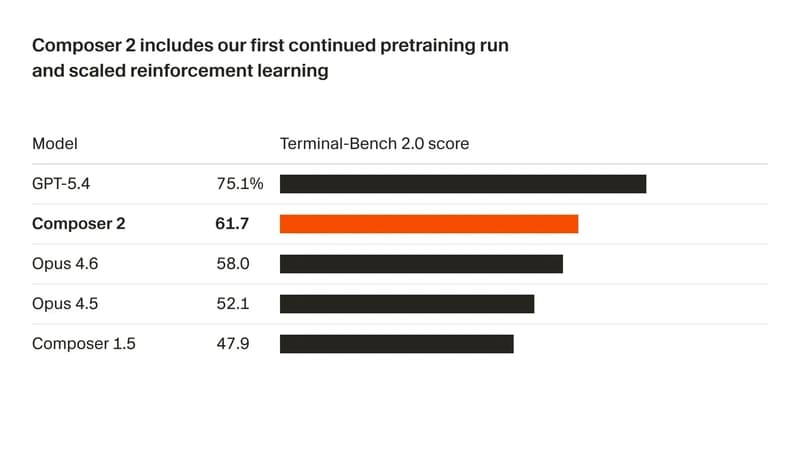
Composer 2: What is new and Compares with Claude Opus 4.6 & GPT-5.4
Dev.to
How UCP Breaks Your E-Commerce Tracking Stack: A Platform-by-Platform Analysis
Dev.to
AI Text Analyzer vs Asking Friends: Which Gives Better Perspective?
Dev.to
[D] Cathie wood claims ai productivity wave is starting, data shows 43% of ceos save 8+ hours weekly
Reddit r/MachineLearning

Microsoft hires top AI researchers from Allen Institute for AI for Suleyman's Superintelligence team
THE DECODER