Segmentation of Gray Matters and White Matters from Brain MRI data
arXiv cs.CV / 4/1/2026
💬 OpinionIdeas & Deep AnalysisModels & Research
Key Points
- The paper presents a modified MedSAM-based foundation model for multi-class segmentation of brain tissues (gray matter and white matter) from MRI data.
- It uses a preprocessing pipeline combining skull stripping (FSL BET) and tissue probability mapping (FSL FAST), then converts volumes into 2D axial/sagittal/coronal slices with multi-class labels.
- The method adapts MedSAM’s mask decoder to predict three classes (background, gray matter, white matter) while freezing the pre-trained image encoder and fine-tuning the prompt encoder and decoder.
- Experiments on the IXI dataset report Dice scores up to 0.8751, indicating strong segmentation performance.
- The authors argue that prompt-based foundation models can be extended to broader and more diverse medical imaging scenarios with relatively small architectural changes.
Related Articles

Show HN: 1-Bit Bonsai, the First Commercially Viable 1-Bit LLMs
Dev.to
I Built an AI Agent That Can Write Its Own Tools When It Gets Stuck
Dev.to

Agent Self-Discovery: How AI Agents Find Their Own Wallets
Dev.to
[P] Federated Adversarial Learning
Reddit r/MachineLearning
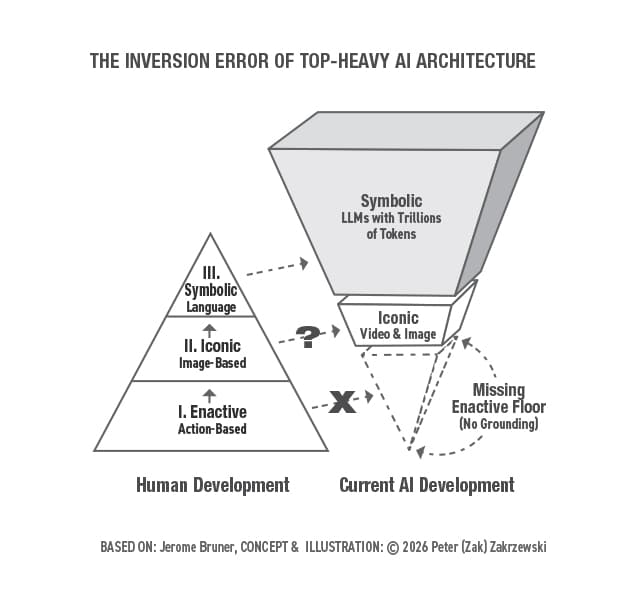
The Inversion Error: Why Safe AGI Requires an Enactive Floor and State-Space Reversibility
Towards Data Science